Hidde DE JONGCBI séminaire publicTitre : Global physiological effects and the analysis of gene regulatory networks
11:00
Natural and synthetic control of growth rate and gene expression in bacteria
Hidde de Jong
Microorganisms adapt the activity of the transcriptional and translational machinery to changes in nutrient availability in their environment. This leads to changes in the expression of a large number of genes, encoding proteins with a variety of cellular functions, and to the adjustment of the growth rate. In recent years, there has been a regained interest in the joint control of gene expression by the activity of the transcriptional and translational machinery and specific transcription factors. I will present a model-based approach to distinguish between these two effects when analyzing changes in gene expression, quantified by means of fluorescent reporter genes. The strength of the approach will be demonstrated by analyzing a genetic circuit involved in the regulation of carbon metabolism in E. coli. I will also present an approach to control the growth rate by putting the transcription of a key component of the gene expression machinery, RNA polymerase, under the control of an inducible promoter. By changing the inducer concentration in the medium, the RNA polymerase concentration can be adjusted and thereby bacterial growth switched between zero and the maximal growth rate supported by the medium. I will show that the proposed synthetic growth switch is a promising tool for gaining a better understanding of bacterial physiology and for applications in synthetic biology and biotechnology.
J. Izard, C. Gomez Balderas, D. Ropers, S. Lacour, X. Song, Y. Yang, A.B. Lindner, J. Geiselmann, H. de Jong. A synthetic growth switch based on controlled expression of RNA polymerase. Molecular Systems Biology, 11(11):840, 2015
V. Zulkower, M. Page, D. Ropers, J. Geiselmann, H. de Jong. Robust reconstruction of gene expression profiles from reporter gene data using linear inversion. Bioinformatics, 31(12):i71-i79, 2015
S. Berthoumieux, H. de Jong, G. Baptist, C. Pinel, C. Ranquet, D. Ropers, and J. Geiselmann. Shared control of gene expression in bacteria by transcription factors and global physiology of the cell. Molecular Systems Biology, 9:634, 2013
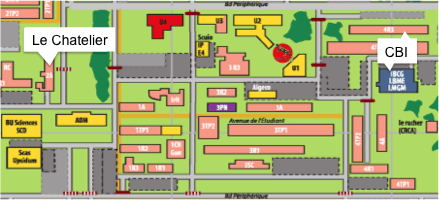
Amphi LE CHATELIER











